-
Single molecular tracking is a valuable approach to investigate the dynamic processes and molecular interactions in soft matter systems, particularly in biological systems. However, understanding the complexity of single molecule motion behaviors in biological systems remains a significant challenge. To address this issue, we propose a two-step classification method based on unsupervised learning to efficiently identify and classify single molecule trajectories. Firstly, we employ an entropy-constrained least square method to distinguish between confined (e.g., immobile) and unconfined diffusion trajectories. Subsequently, statistical tests are utilized to categorize the unconfined trajectories into different diffusion modes such as sub-diffusion, normal diffusion, and super-diffusion. By applying this method, we analyze the diffusion motion of single molecules in both DOPC model cell membranes and living cell membranes while uncovering their distinct responses to cholesterol composition. Our findings demonstrate that both model membranes and living cell membranes exhibit diverse molecular diffusion modes. Specifically, in the DOPC model membrane system, the presence of cholesterol components impedes lipid diffusion within the membrane. The degree of inhibition is positively correlated with the amount of cholesterol present. For instance, as the cholesterol content in the membrane increases from 0 to 20% (DOPC:Chol = 4∶1) and 50% (DOPC:Chol = 1∶1), there is an increase in the proportion of molecules, exhibiting confined diffusion and sub-diffusion (from 55% to 45%), while there is a decrease in the proportion of molecules, displaying normal diffusion and super-diffusion (from 45% to 35%). The ensemble diffusion coefficient of molecules in the membrane significantly decreases, which can be attributed to both a decrease in velocity among fast-moving molecules. Interestingly, after using MeβCD to remove cholesterol, the single-molecule mobility within the DOPC/Chol composite membrane system is restored to a level similar to that of the pure DOPC membrane. Conversely, in the living cell membrane system, the diffusion coefficient values of molecules are significantly lower than those observed in the model membrane system; furthermore, the removal of cholesterol further slows down the molecular diffusion rate. This study contributes to understanding the intricacies of biomolecular motility and its dependence on environmental factors from a perspective of single molecular motion. -
Keywords:
- single molecular motion /
- unsupervised learning /
- model cell membrane /
- cholesterol
[1] Jacobson K, Liu P, Lagerholm B C 2019 Cell 177 806
 Google Scholar
Google Scholar
[2] He W, Song H, Su Y, et al. 2016 Nat. Commun. 7 11701
 Google Scholar
Google Scholar
[3] Golan Y, Sherman E 2017 Nat. Commun. 8 15851
 Google Scholar
Google Scholar
[4] Subczynski W K, Pasenkiewicz-Gierula M, Widomska J, Mainali L, Raguz M 2017 Cell Biochem. Biophys. 75 369
 Google Scholar
Google Scholar
[5] van Meer G, Voelker D R, Feigenson G W 2008 Nat. Rev. Mol. Cell Biol. 9 112
 Google Scholar
Google Scholar
[6] Liu Y, Zheng X, Guan D, Jiang X, Hu G 2022 ACS Nano 16 16054
 Google Scholar
Google Scholar
[7] Lyman E 2021 Biophys. J. 120 1777
 Google Scholar
Google Scholar
[8] Zhang X, Barraza K M, Beauchamp J L 2018 P. Natl. Acad. Sci. USA 115 3255
 Google Scholar
Google Scholar
[9] Chakraborty S, Doktorova M, Molugu T R, et al. 2020 P. Natl. Acad. Sci. USA 117 21896
 Google Scholar
Google Scholar
[10] Pohnl M, Trollmann M F W, Bockmann R A 2023 Nat. Commun. 14 8038
 Google Scholar
Google Scholar
[11] Fernandez-Perez E J, Sepulveda F J, Peters C, et al. 2018 Front. Aging Neurosci. 10 226
 Google Scholar
Google Scholar
[12] Doole F T, Kumarage T, Ashkar R, Brown M F 2022 J. Membr. Biol. 255 385
 Google Scholar
Google Scholar
[13] Byfield F J, Aranda-Espinoza H, Romanenko V G, Rothblat G H, Levitan I 2004 Biophys. J. 87 3336
 Google Scholar
Google Scholar
[14] Yang S T, Kreutzberger A J B, Lee J, Kiessling V, Tamm L K 2016 Chem. Phys. Lipids 199 136
 Google Scholar
Google Scholar
[15] Norregaard K, Metzler R, Ritter C M, Berg-Sorensen K, Oddershede L B 2017 Chem. Rev. 117 4342
 Google Scholar
Google Scholar
[16] Ge F, Du Y, He Y 2022 ACS Nano 16 5325
 Google Scholar
Google Scholar
[17] Chen P Y, Yue H, Zhai X B, Huang Z H, Ma G H, Wei W, Yan L T 2019 Sci. Adv. 5 eaaw3192
 Google Scholar
Google Scholar
[18] Jeon J H, Javanainen M, Martinez-Seara H, Metzler R, Vattulainen I 2016 Phys. Rev. X 6 021006
 Google Scholar
Google Scholar
[19] Xu C, Yang K, Yuan B 2023 J. Phys. Chem. Lett. 14 854
 Google Scholar
Google Scholar
[20] Xu C, Ma W, Wang K, He K, Chen Z, Liu J, Yang K, Yuan B 2020 J. Phys. Chem. Lett. 11 4834
 Google Scholar
Google Scholar
[21] Pinholt H D, Bohr S S R, Iversen J F, Boomsma W, Hatzakis N S 2021 P. Natl. Acad. Sci. USA 118 e2104624118
 Google Scholar
Google Scholar
[22] Muñoz-Gil G, Garcia-March M A, Manzo C, Martín-Guerrero J D, Lewenstein M 2020 New J. Phys. 22 013010
 Google Scholar
Google Scholar
[23] Cherstvy A G, Thapa S, Wagner C E, Metzler R 2019 Soft Matter 15 2526
 Google Scholar
Google Scholar
[24] Granik N, Weiss L E, Nehme E, Levin M, Chein M, Perlson E, Roichman Y, Shechtman Y 2019 Biophys. J. 117 185
 Google Scholar
Google Scholar
[25] Janczura J, Kowalek P, Loch-Olszewska H, Szwabinski J, Weron A 2020 Phys. Rev. E 102 032402
 Google Scholar
Google Scholar
[26] Barkai E, Garini Y, Metzler R 2012 Phys. Today 65 29
 Google Scholar
Google Scholar
[27] Krapf D, Metzler R 2019 Phys. Today 72 48
 Google Scholar
Google Scholar
[28] Wu J F, Xu C, Ye Z F, Chen H B, Wang Y P, Yang K, Yuan B 2023 Small 19 2301713
 Google Scholar
Google Scholar
[29] Yamamoto E, Akimoto T, Kalli A C, Yasuoka K, Sansom M S P 2017 Sci. Adv. 3 e1601871
 Google Scholar
Google Scholar
[30] Feder T J, Brust-Mascher I, Slattery J P, Baird B, Webb W W 1996 Biophys. J. 70 2767
 Google Scholar
Google Scholar
[31] Briane V, Kervrann C, Vimond M 2018 Phys. Rev. E 97 062121
 Google Scholar
Google Scholar
[32] Lanoiselée Y, Sikora G, Grzesiek A, Grebenkov D S, Wylomanska A 2018 Phys. Rev. E 98 062139
 Google Scholar
Google Scholar
[33] Sikora G, Teuerle M, Wylomanska A, Grebenkov D 2017 Phys. Rev. E 96 022132
 Google Scholar
Google Scholar
[34] Saxton M J, Jacobson K 1997 Annu. Rev. Biophys. Biomol. Struct. 26 373
 Google Scholar
Google Scholar
[35] Kusumi A, Sako Y, Yamamoto M 1993 Biophys. J. 65 2021
 Google Scholar
Google Scholar
[36] Saxton M J 1993 Biophys. J. 64 1766
 Google Scholar
Google Scholar
[37] Shannon C E 1948 Bell Syst. Tech. J. 27 379
 Google Scholar
Google Scholar
[38] Wright S J, Tenny M J 2004 Siam J. Optim. 14 1074
 Google Scholar
Google Scholar
[39] Raja M A Z, Ahmed U, Zameer A, Kiani A K, Chaudhary N I 2019 Neural. Comput. Appl. 31 447
 Google Scholar
Google Scholar
[40] Zhang Y, Yao F, Iu H H C, Fernando T, Wong K P 2013 J. Mod. Power Syst. Clean Energy 1 231
 Google Scholar
Google Scholar
[41] Weron A, Janczura J, Boryczka E, Sungkaworn T, Calebiro D 2019 Phys. Rev. E 99 042149
 Google Scholar
Google Scholar
[42] Hubicka K, Janczura J 2020 Phys. Rev. E 101 022107
 Google Scholar
Google Scholar
[43] Xu R, Zhang W T, Jin T, Tu W, Xu C, Wei Y, Han W, Yang K, Yuan B 2024 ACS Appl. Mater. Interfaces 16 6813
 Google Scholar
Google Scholar
[44] Li L, Ji J, Song F, Hu J 2023 J. Mol. Biol. 435 167787
 Google Scholar
Google Scholar
[45] Li L, Hou R H, Shi X H, et al. 2024 Commun. Phys. 7 174
 Google Scholar
Google Scholar
[46] Gao J, Hou R, Li L, Hu J 2021 Front. Mol. Biosci. 8 811711
 Google Scholar
Google Scholar
[47] 陆越, 马建兵, 滕翠娟, 陆颖, 李明, 徐春华 2018 67 088201
 Google Scholar
Google Scholar
Lu Y, Ma J B, Teng C J, Lu Y, Li M, Xu C H 2018 Acta Phys. Sin. 67 088201
 Google Scholar
Google Scholar
[48] Gao J, Hou R, Hu W, et al. 2024 J. Phys. Chem. B 128 4735
 Google Scholar
Google Scholar
-
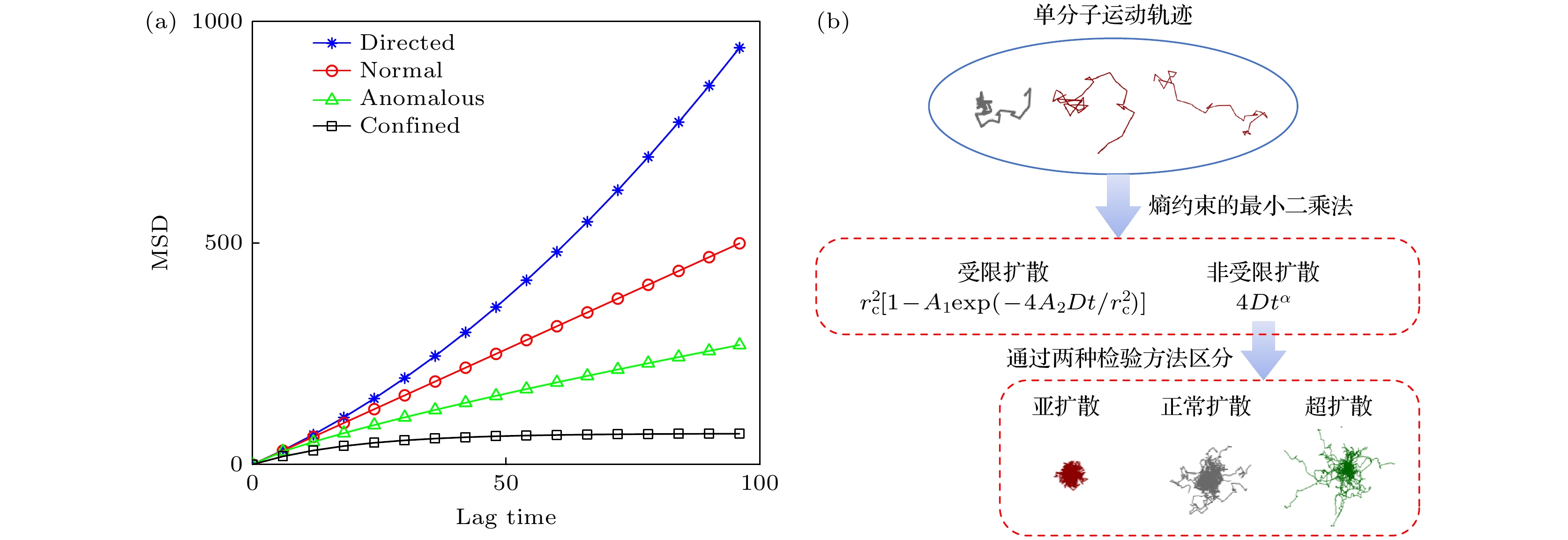
图 1 单分子扩散运动的不同模式 (a)模拟运动模型的MSD曲线, 即定向扩散(蓝色)、正常扩散(红色)、异常扩散(绿色)和受限扩散(黑色), 横坐标为滞后时间(lag time); (b)采用“两步归类法”对单分子轨迹进行识别和分类的步骤, 第一步区分受限扩散与非受限扩散, 第二步将非受限扩散进一步区分
Figure 1. Different modes of single-molecule diffusion. (a) Simulated mean square displacement (MSD) curves depicting directed diffusion (blue), normal diffusion (red), anomalous diffusion (green), and confined diffusion (black). (b) The “two-step classification method” employed to identify and classify single-molecule trajectories: the first step involves distinguishing between confined and unconfined diffusion, followed by further differentiation of unconfined diffusion in the second step.
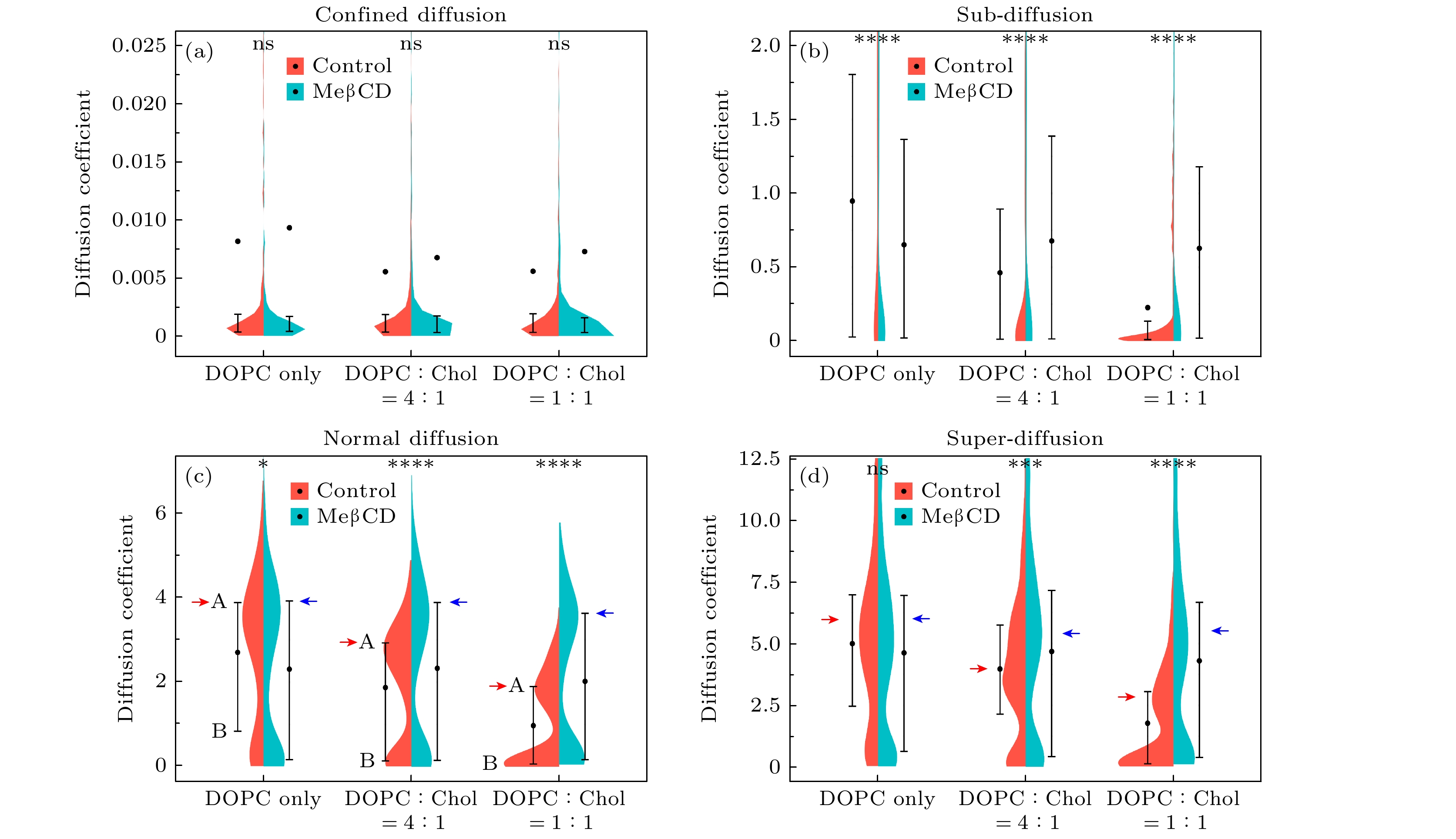
图 2 不同模型膜体系内单分子扩散运动分析, 即利用“两步归类法”将其分为受限扩散(Confined diffusion)、亚扩散(Sub-diffusion)、正常扩散(Normal diffusion)、超扩散(Super-diffusion) 4个子群, 分别计算其单分子轨迹的时间平均MSD并作出其分布. 轨迹数目见表1
Figure 2. Analysis of single lipid diffusion in various model membrane systems using the “two-step classification method”. Trajectories were categorized into four subgroups, namely confined diffusion, sub-diffusion, normal diffusion, and super-diffusion. The time-averaged MSD of individual trajectories was calculated to obtain the distribution. The number of trajectories is presented in Table 1.
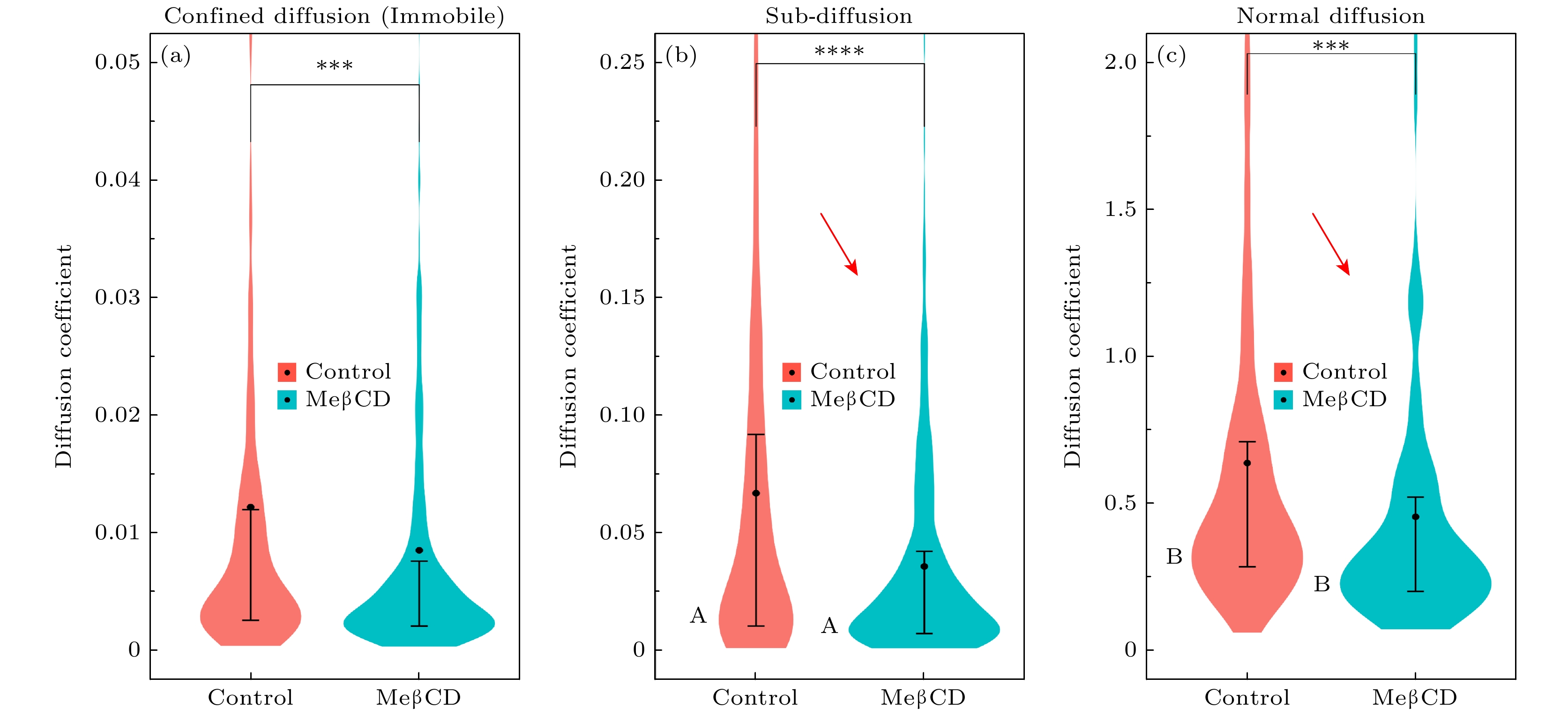
图 3 B16活细胞膜(MeβCD处理前或处理后)内单分子扩散运动分析, 即利用“两步归类法”将其分为受限扩散(Confined diffusion)、亚扩散(Sub-diffusion)、正常扩散(Normal diffusion) 3个子群, 分别计算其单分子轨迹的时间平均MSD并作出其分布, 轨迹数目见表2
Figure 3. Analysis of single-molecule diffusion in live cell membranes (before or after MeβCD treatment). This analysis employs a two-step classification method to divide the data into subgroups of confined diffusion, sub-diffusion, and normal diffusion. The time-averaged MSD is calculated for individual trajectories and its distribution is plotted. The number of trajectories analyzed is provided in Table 2.
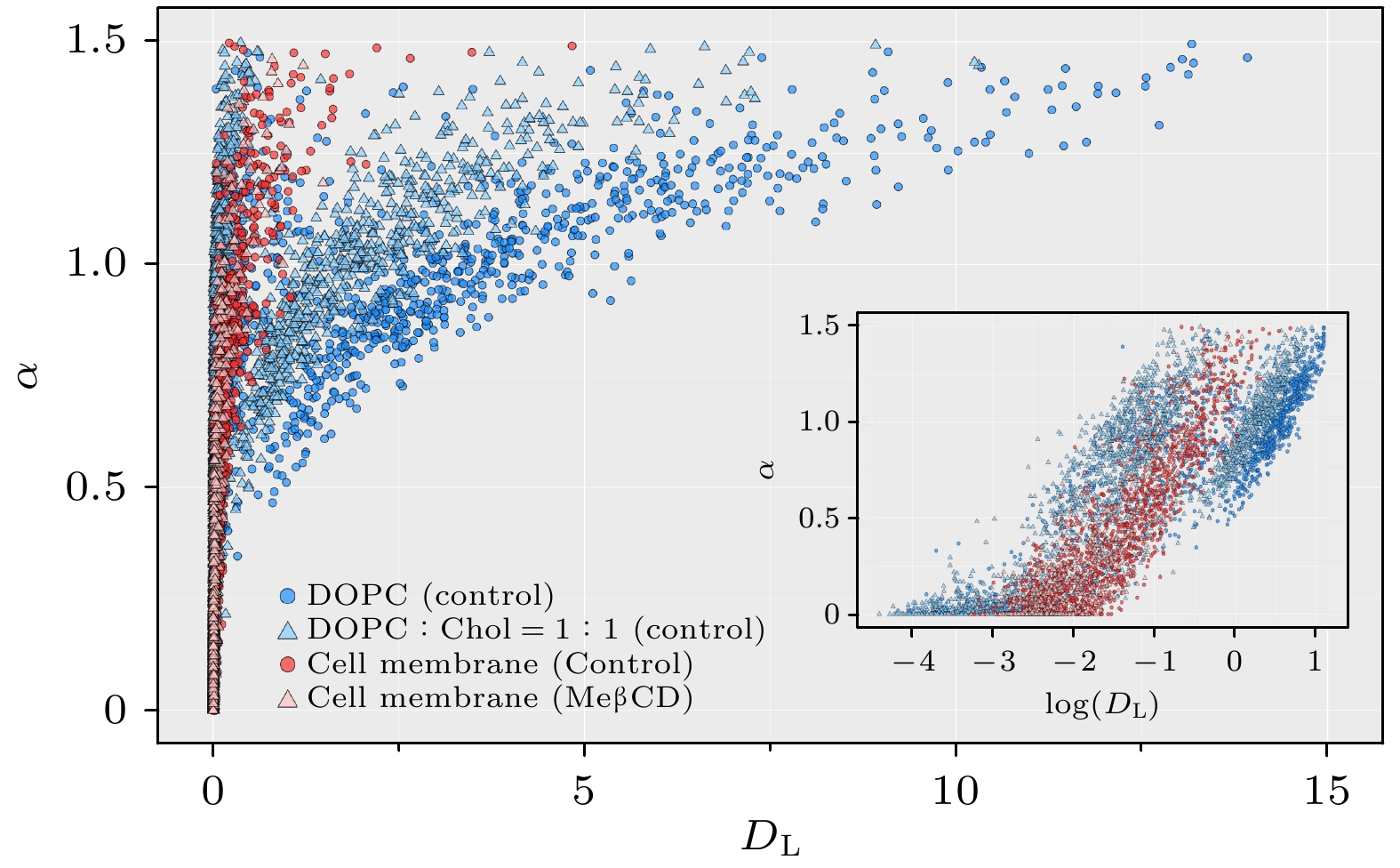
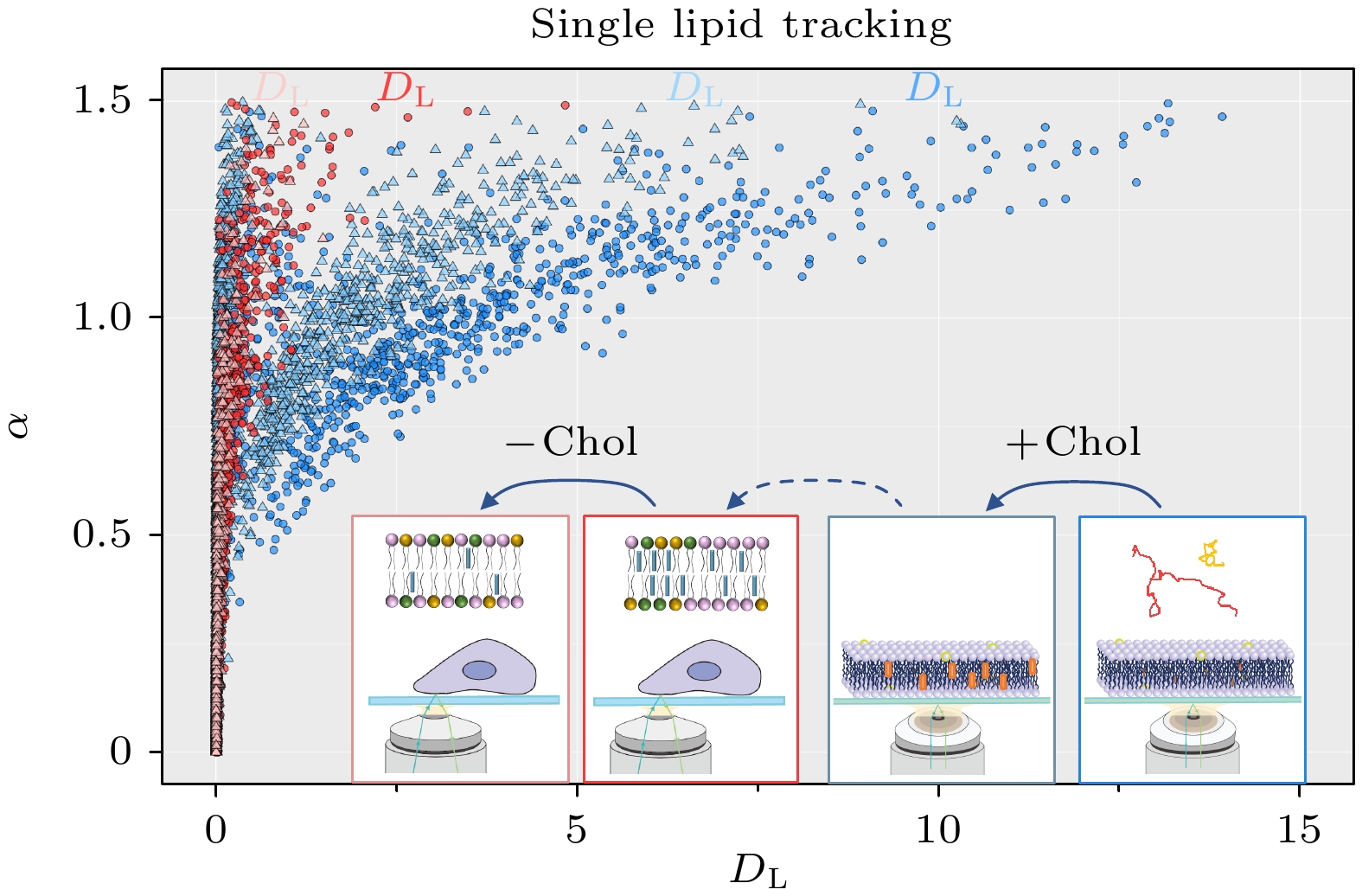
图 4 不同体系内DL和α值分布, 包括纯DOPC膜(包含1324条轨迹)、DOPC∶Chol = 1∶1混合膜(2347条轨迹)、活细胞膜经MeβCD处理之前(1436条轨迹)和之后(784条轨迹), 其中插图中横坐标以对数形式显示
Figure 4. Distribution of DL and α values in different systems: pure DOPC membrane (1324 trajectories), DOPC∶Chol = 1∶1 mixed membrane (2347 trajectories), living cell membrane before MeβCD treatment (1436 trajectories), and after MeβCD treatment (784 trajectories). The horizontal coordinate is presented logarithmically in the inset.
表 1 不同模型膜体系内单脂质分子轨迹分类及相应的系综扩散系数和异常指数数值, 其中轨迹长度为60帧
Table 1. Classification of single lipid trajectories in different model membrane systems and corresponding ensemble diffusion coefficient and anomaly index values. The trajectory length is 60 frames.
Subgroup Control +MeβCD DOPC DOPC:CHOL
= 4∶1DOPC:CHOL
= 1∶1DOPC DOPC:CHOL
= 4∶1DOPC:CHOL
= 1∶1Trajectory
numberConfined 282 383 497 293 334 330 Sub-diffusion 943 1143 1036 947 799 811 Normal diffusion 318 294 290 305 200 207 Super-diffusion 667 577 524 647 448 449 Proportion/% Confined 12.76 15.98 21.18 13.37 18.75 18.36 Sub-diffusion 42.67 47.68 44.14 43.20 44.86 45.13 Normal diffusion 14.39 12.27 12.36 13.91 11.23 11.52 Super-diffusion 30.18 24.07 22.33 29.52 25.15 24.99 Diffusion coefficient
DL/(μm2·s–1)Confined 0.045 0.032 0.029 0.064 0.034 0.061 Sub-diffusion 1.086 0.582 0.308 0.805 0.784 0.717 Normal diffusion 2.421 1.581 0.870 2.100 2.051 1.781 Super-diffusion 4.149 3.211 1.444 3.908 3.650 3.381 Anomaly
index (α)Confined 0.116 0.094 0.068 0.207 0.099 0.171 Sub-diffusion 0.709 0.629 0.512 0.629 0.635 0.614 Normal diffusion 0.909 0.869 0.916 0.948 0.910 0.874 Super-diffusion 1.133 1.103 1.178 1.141 1.132 1.132 表 2 经MeβCD去除胆固醇前后活细胞膜内单分子轨迹分类及相应的系综扩散系数和异常指数数值, 其中轨迹长度为20帧
Table 2. Classification of single-molecule trajectory in living cell membranes before and after cholesterol depletion by MeβCD, along with corresponding ensemble-averaged diffusion coefficient and anomaly index values. The trajectory length is 20 frames.
Subgroup Control +MeβCD Trajectory
numberConfined 452 304 Sub-diffusion 645 377 Normal diffusion 339 103 Proportion/% Confined 31.48 38.78 Sub-diffusion 44.92 48.09 Normal diffusion 23.61 13.14 Diffusion coefficient
DL/(μm2·s–1)Confined 0.017 0.011 Sub-diffusion 0.069 0.040 Normal diffusion 0.465 0.260 Anomaly
index (α)Confined 0.146 0.095 Sub-diffusion 0.437 0.461 Normal diffusion 0.983 0.861 -
[1] Jacobson K, Liu P, Lagerholm B C 2019 Cell 177 806
 Google Scholar
Google Scholar
[2] He W, Song H, Su Y, et al. 2016 Nat. Commun. 7 11701
 Google Scholar
Google Scholar
[3] Golan Y, Sherman E 2017 Nat. Commun. 8 15851
 Google Scholar
Google Scholar
[4] Subczynski W K, Pasenkiewicz-Gierula M, Widomska J, Mainali L, Raguz M 2017 Cell Biochem. Biophys. 75 369
 Google Scholar
Google Scholar
[5] van Meer G, Voelker D R, Feigenson G W 2008 Nat. Rev. Mol. Cell Biol. 9 112
 Google Scholar
Google Scholar
[6] Liu Y, Zheng X, Guan D, Jiang X, Hu G 2022 ACS Nano 16 16054
 Google Scholar
Google Scholar
[7] Lyman E 2021 Biophys. J. 120 1777
 Google Scholar
Google Scholar
[8] Zhang X, Barraza K M, Beauchamp J L 2018 P. Natl. Acad. Sci. USA 115 3255
 Google Scholar
Google Scholar
[9] Chakraborty S, Doktorova M, Molugu T R, et al. 2020 P. Natl. Acad. Sci. USA 117 21896
 Google Scholar
Google Scholar
[10] Pohnl M, Trollmann M F W, Bockmann R A 2023 Nat. Commun. 14 8038
 Google Scholar
Google Scholar
[11] Fernandez-Perez E J, Sepulveda F J, Peters C, et al. 2018 Front. Aging Neurosci. 10 226
 Google Scholar
Google Scholar
[12] Doole F T, Kumarage T, Ashkar R, Brown M F 2022 J. Membr. Biol. 255 385
 Google Scholar
Google Scholar
[13] Byfield F J, Aranda-Espinoza H, Romanenko V G, Rothblat G H, Levitan I 2004 Biophys. J. 87 3336
 Google Scholar
Google Scholar
[14] Yang S T, Kreutzberger A J B, Lee J, Kiessling V, Tamm L K 2016 Chem. Phys. Lipids 199 136
 Google Scholar
Google Scholar
[15] Norregaard K, Metzler R, Ritter C M, Berg-Sorensen K, Oddershede L B 2017 Chem. Rev. 117 4342
 Google Scholar
Google Scholar
[16] Ge F, Du Y, He Y 2022 ACS Nano 16 5325
 Google Scholar
Google Scholar
[17] Chen P Y, Yue H, Zhai X B, Huang Z H, Ma G H, Wei W, Yan L T 2019 Sci. Adv. 5 eaaw3192
 Google Scholar
Google Scholar
[18] Jeon J H, Javanainen M, Martinez-Seara H, Metzler R, Vattulainen I 2016 Phys. Rev. X 6 021006
 Google Scholar
Google Scholar
[19] Xu C, Yang K, Yuan B 2023 J. Phys. Chem. Lett. 14 854
 Google Scholar
Google Scholar
[20] Xu C, Ma W, Wang K, He K, Chen Z, Liu J, Yang K, Yuan B 2020 J. Phys. Chem. Lett. 11 4834
 Google Scholar
Google Scholar
[21] Pinholt H D, Bohr S S R, Iversen J F, Boomsma W, Hatzakis N S 2021 P. Natl. Acad. Sci. USA 118 e2104624118
 Google Scholar
Google Scholar
[22] Muñoz-Gil G, Garcia-March M A, Manzo C, Martín-Guerrero J D, Lewenstein M 2020 New J. Phys. 22 013010
 Google Scholar
Google Scholar
[23] Cherstvy A G, Thapa S, Wagner C E, Metzler R 2019 Soft Matter 15 2526
 Google Scholar
Google Scholar
[24] Granik N, Weiss L E, Nehme E, Levin M, Chein M, Perlson E, Roichman Y, Shechtman Y 2019 Biophys. J. 117 185
 Google Scholar
Google Scholar
[25] Janczura J, Kowalek P, Loch-Olszewska H, Szwabinski J, Weron A 2020 Phys. Rev. E 102 032402
 Google Scholar
Google Scholar
[26] Barkai E, Garini Y, Metzler R 2012 Phys. Today 65 29
 Google Scholar
Google Scholar
[27] Krapf D, Metzler R 2019 Phys. Today 72 48
 Google Scholar
Google Scholar
[28] Wu J F, Xu C, Ye Z F, Chen H B, Wang Y P, Yang K, Yuan B 2023 Small 19 2301713
 Google Scholar
Google Scholar
[29] Yamamoto E, Akimoto T, Kalli A C, Yasuoka K, Sansom M S P 2017 Sci. Adv. 3 e1601871
 Google Scholar
Google Scholar
[30] Feder T J, Brust-Mascher I, Slattery J P, Baird B, Webb W W 1996 Biophys. J. 70 2767
 Google Scholar
Google Scholar
[31] Briane V, Kervrann C, Vimond M 2018 Phys. Rev. E 97 062121
 Google Scholar
Google Scholar
[32] Lanoiselée Y, Sikora G, Grzesiek A, Grebenkov D S, Wylomanska A 2018 Phys. Rev. E 98 062139
 Google Scholar
Google Scholar
[33] Sikora G, Teuerle M, Wylomanska A, Grebenkov D 2017 Phys. Rev. E 96 022132
 Google Scholar
Google Scholar
[34] Saxton M J, Jacobson K 1997 Annu. Rev. Biophys. Biomol. Struct. 26 373
 Google Scholar
Google Scholar
[35] Kusumi A, Sako Y, Yamamoto M 1993 Biophys. J. 65 2021
 Google Scholar
Google Scholar
[36] Saxton M J 1993 Biophys. J. 64 1766
 Google Scholar
Google Scholar
[37] Shannon C E 1948 Bell Syst. Tech. J. 27 379
 Google Scholar
Google Scholar
[38] Wright S J, Tenny M J 2004 Siam J. Optim. 14 1074
 Google Scholar
Google Scholar
[39] Raja M A Z, Ahmed U, Zameer A, Kiani A K, Chaudhary N I 2019 Neural. Comput. Appl. 31 447
 Google Scholar
Google Scholar
[40] Zhang Y, Yao F, Iu H H C, Fernando T, Wong K P 2013 J. Mod. Power Syst. Clean Energy 1 231
 Google Scholar
Google Scholar
[41] Weron A, Janczura J, Boryczka E, Sungkaworn T, Calebiro D 2019 Phys. Rev. E 99 042149
 Google Scholar
Google Scholar
[42] Hubicka K, Janczura J 2020 Phys. Rev. E 101 022107
 Google Scholar
Google Scholar
[43] Xu R, Zhang W T, Jin T, Tu W, Xu C, Wei Y, Han W, Yang K, Yuan B 2024 ACS Appl. Mater. Interfaces 16 6813
 Google Scholar
Google Scholar
[44] Li L, Ji J, Song F, Hu J 2023 J. Mol. Biol. 435 167787
 Google Scholar
Google Scholar
[45] Li L, Hou R H, Shi X H, et al. 2024 Commun. Phys. 7 174
 Google Scholar
Google Scholar
[46] Gao J, Hou R, Li L, Hu J 2021 Front. Mol. Biosci. 8 811711
 Google Scholar
Google Scholar
[47] 陆越, 马建兵, 滕翠娟, 陆颖, 李明, 徐春华 2018 67 088201
 Google Scholar
Google Scholar
Lu Y, Ma J B, Teng C J, Lu Y, Li M, Xu C H 2018 Acta Phys. Sin. 67 088201
 Google Scholar
Google Scholar
[48] Gao J, Hou R, Hu W, et al. 2024 J. Phys. Chem. B 128 4735
 Google Scholar
Google Scholar
-
 2024年73卷188702补充材料.pdf
2024年73卷188702补充材料.pdf

 2024年73卷188702补充材料-模型膜和活细胞膜内单分子运动轨迹的扩散系数、异常指数、位移的三维分布视频.gif
2024年73卷188702补充材料-模型膜和活细胞膜内单分子运动轨迹的扩散系数、异常指数、位移的三维分布视频.gif

Catalog
Metrics
- Abstract views: 4367
- PDF Downloads: 123
- Cited By: 0














 DownLoad:
DownLoad: