-
Transcription termination is a critical step for gene regulation and genome integrity among all kingdoms of life. In Saccharomyces cerevisiae, one of the major termination pathways is accomplished by Sen1 helicase, a homolog to human Senataxin (SETX), defection of which raises the diseases for the central nervus system of human. Although it has been proposed that Sen1 translocates along nucleic acids by consuming adenosine triphosphates (ATPs) during termination, the mechanism for this translocation activity of Sen1 has not been well understood. In this work, our aim is to investigate the mechanism of Sen1 translocation by measuring the interactions between Sen1 and different types of nucleic acids by polyacrylamide gel electrophoresis (PAGE) assay or single-molecule Fӧrster resonance energy transfer (FRET) assay. We firstly observe the unwinding activity of Sen1 on a tailed duplex DNA in the presence of 1 mM ATP via PAGE assay, where the translocation activity of Sen1 is involved. As the binding activity is crucial for translocation, then we examine the binding affinity of Sen1 to the single-stranded DNA via PAGE assay, revealing a stable binding of Sen1 with an occupied length of nucleic acids of less than 24 nt. In the presence of 1 µM ATP, we observe that Sen1 dynamically binds to and dissociates from the tailed duplex DNA in the single-molecule FRET assay. By titrating ATP concentrations from 1–500 µM, we observe a gradual decrease in the mean durations of Sen1 binding, suggesting an ATP-dependent binding affinity of Sen1 to single-stranded DNA. We then fit these mean durations to the classical Michaelis-Menten model and obtain a minimum binding duration of (0.18 ± 0.01) s at saturating ATP concentrations and Km of (13.1 ± 0.1) µM for the ATP-dependent binding of Sen1. This result is consistent with that from a translocation activity of Sen1. Taking into account the translocation length of the half of the single-stranded tail, i.e. 13 nt, a mean rate of 70 nt/s is estimated. Reversing the translocation direction, we observe an increase in the duration of Sen1 binding to the single-stranded tail, which suggests an impediment of DNA duplex in front of Sen1 translocation or the possible duplex DNA unwinding activity of Sen1. Our quantitative measurements on Sen1 translocation are helpful in deepening our understanding of the mechanism of eukaryotic transcription termination by Sen1.
-
Keywords:
- Sen1 helicase /
- translocation /
- single-molecule Fӧrster resonance energy transfer /
- transcription termination
[1] Roeder R G 2019 Nat. Struct. Mol. Biol. 26 783
 Google Scholar
Google Scholar
[2] Jonkers I, Lis J T 2015 Nat. Rev. Mol. Cell. Biol. 16 167
 Google Scholar
Google Scholar
[3] Abbondanzieri E A, Greenleaf W J, Shaevitz J W, Landick R, Block S M 2005 Nature 438 460
 Google Scholar
Google Scholar
[4] You L, Omollo E O, Yu C, Mooney R A, Shi J, Shen L, Wu X, Wen A, He D, Zeng Y, Feng Y, Landick R, Zhang Y 2023 Nature 613 783
 Google Scholar
Google Scholar
[5] Molodtsov V, Wang C, Firlar E, Kaelber J T, Ebright R H 2023 Nature 614 367
 Google Scholar
Google Scholar
[6] West S, Gromak N, Proudfoot N J 2004 Nature 432 522
 Google Scholar
Google Scholar
[7] Porrua O, Libri D 2013 Nat. Struct. Mol. Biol. 20 884
 Google Scholar
Google Scholar
[8] Wang S, Han Z, Libri D, Porrua O, Strick T R 2019 Nat. Commun. 10 1545
 Google Scholar
Google Scholar
[9] Vasiljeva L, Kim M, Mutschler H, Buratowski S, Meinhart A 2008 Nat. Struct. Mol. Biol. 15 795
 Google Scholar
Google Scholar
[10] Arndt K M, Reines D 2015 Annu. Rev. Biochem. 84 381
 Google Scholar
Google Scholar
[11] Rondón A G, Mischo H E, Kawauchi J, Proudfoot N J 2009 Mol. Cell. 36 88
 Google Scholar
Google Scholar
[12] Jia X, Li Y, Wang T, Bi L, Guo L, Chen Z, Zhang X, Ye S, Chen J, Yang B, Sun B 2023 Embo. J. 42 e111703
 Google Scholar
Google Scholar
[13] Saper G, Hess H 2020 Chem. Rev. 120 288
 Google Scholar
Google Scholar
[14] Nishizaka T 2010 Adv. Biochem. Eng. Biotechnol. 119 3
 Google Scholar
Google Scholar
[15] Martin-Tumasz S, Brow D A 2015 J. Biol. Chem. 290 22880
 Google Scholar
Google Scholar
[16] Han Z, Libri D, Porrua O 2017 Nucleic. Acids. Res. 45 1355
 Google Scholar
Google Scholar
[17] Skourti-Stathaki K, Proudfoot N J, Gromak N 2011 Mol. Cell. 42 794
 Google Scholar
Google Scholar
[18] Ha T, Enderle T, Ogletree D F, Chemla D S, Selvin P R, Weiss S 1996 Proc. Natl. Acad. Sci. USA 93 6264
 Google Scholar
Google Scholar
[19] Maki A H, Co T 1976 Biochemistry 15 1229
 Google Scholar
Google Scholar
[20] Leonaitė B, Han Z, Basquin J, Bonneau F, Libri D, Porrua O, Conti E 2017 Embo. J. 36 1590
 Google Scholar
Google Scholar
[21] Shi J, Wang F, Li F, Wang L, Xiong Y, Wen A, Jin Y, Jin S, Gao F, Feng Z, Li J, Zhang Y, Shang Z, Wang S, Feng Y, Lin W 2022 Nucleic. Acids. Res. 50 5974
 Google Scholar
Google Scholar
[22] Fiorini F, Bagchi D, Le Hir H, Croquette V 2015 Nat. Commun. 6 7581
 Google Scholar
Google Scholar
[23] Chakrabarti S, Jayachandran U, Bonneau F, Fiorini F, Basquin C, Domcke S, Le Hir H, Conti E 2011 Mol. Cell. 41 693
 Google Scholar
Google Scholar
[24] Hazelbaker D Z, Marquardt S, Wlotzka W, Buratowski S 2013 Mol. Cell. 49 55
 Google Scholar
Google Scholar
-
图 1 聚丙烯酰胺凝胶电泳表征Sen1 HD解旋双链DNA活性 (a) Sen1解旋酶蛋白结构域模式图. (b) 实验方法示意图, Sen1 HD作用于5'-50ss-20duplex底物, 通过水解ATP获得能量来解开20-nt FAM ssDNA. (c) 对FAM荧光成像结果1道为20-nt FAM ssDNA的条带位置; 2道为5'-50ss-20duplex的条带位置; 3道为5'-50ss-20duplex与20-nt竞争ssDNA 30 ℃孵育20 min的结果, 无20-nt FAM ssDNA产生; 4道为5'-50ss-20duplex、20-nt竞争ssDNA和Sen1 HD 30 ℃孵育20 min的结果, 无20-nt FAM ssDNA产生; 5道为5'-50ss-20duplex, 20-nt竞争ssDNA, Sen1 HD, ATP条件下, 30 ℃孵育20 min的结果, 产生20-nt FAM ssDNA
Figure 1. Characterization of Sen1 HD unwinding activity on double-stranded DNA via PAGE assay. (a) Domain pattern of Sen1. (b) Schematic of PAGE assay, Sen1 HD acts on the 5'-50ss-20duplex substrate to unwind the 20-nt FAM ssDNA by hydrolyzing ATP. (c) Fluorescence imaging of PAGE result: the first lane represents the band position of 20-nt FAM ssDNA; the second lane represents the band position of 5'-50ss-20duplex; the third lane represented 5'-50ss-20duplex with 20-nt competed ssDNA incubated at 30 ℃ for 20 min, but no 20-nt FAM ssDNA was produced; the fourth lane represented 5'-50ss-20duplex with 20-nt competed ssDNA and Sen1 HD incubated at 30 ℃ for 20 min, but no 20-nt FAM ssDNA was produced; the fifth lane, 20-nt FAM ssDNA was produced after incubation for 20 min at 30 ℃ under the condition of 5'-50ss-20duplex, 20-nt competed ssDNA, Sen1 HD and ATP.
图 2 Sen1 HD与Cy3-48ssDNA结合的PAGE实验 (a) Sen1 HD与Cy3-48ssDNA结合示意图. (b) Cy3荧光成像结果. 第1道为Cy3-48ssDNA原始长度; 第2道为5 nM Sen1 HD与Cy3-48ssDNA 30 ℃孵育30 min的结果; 第3道为25 nM Sen1 HD与Cy3-48ssDNA 30 ℃孵育30 min的结果; 第4道为50 nM Sen1 HD与Cy3-48ssDNA 30 ℃孵育30 min的结果
Figure 2. Sen1 HD binding activity to Cy3-48ssDNA characterized via PAGE assay. (a) Schematic of Sen1 HD binding to Cy3-48ssDNA. (b) Fluorescence imaging of Cy3 fluorophore: the first lane represents the original length of Cy3-48ssDNA; the second lane represents the result of 5 nM Sen1 HD incubation with Cy3-48ssDNA at 30 ℃ for 30 min; the third lane represents the result of 25 nM Sen1 HD incubation with Cy3-48ssDNA at 30 ℃ for 30 minutes; the fourth lane represents the result of 50 nM Sen1 HD incubation with Cy3-48ssDNA at 30 ℃ for 30 min.
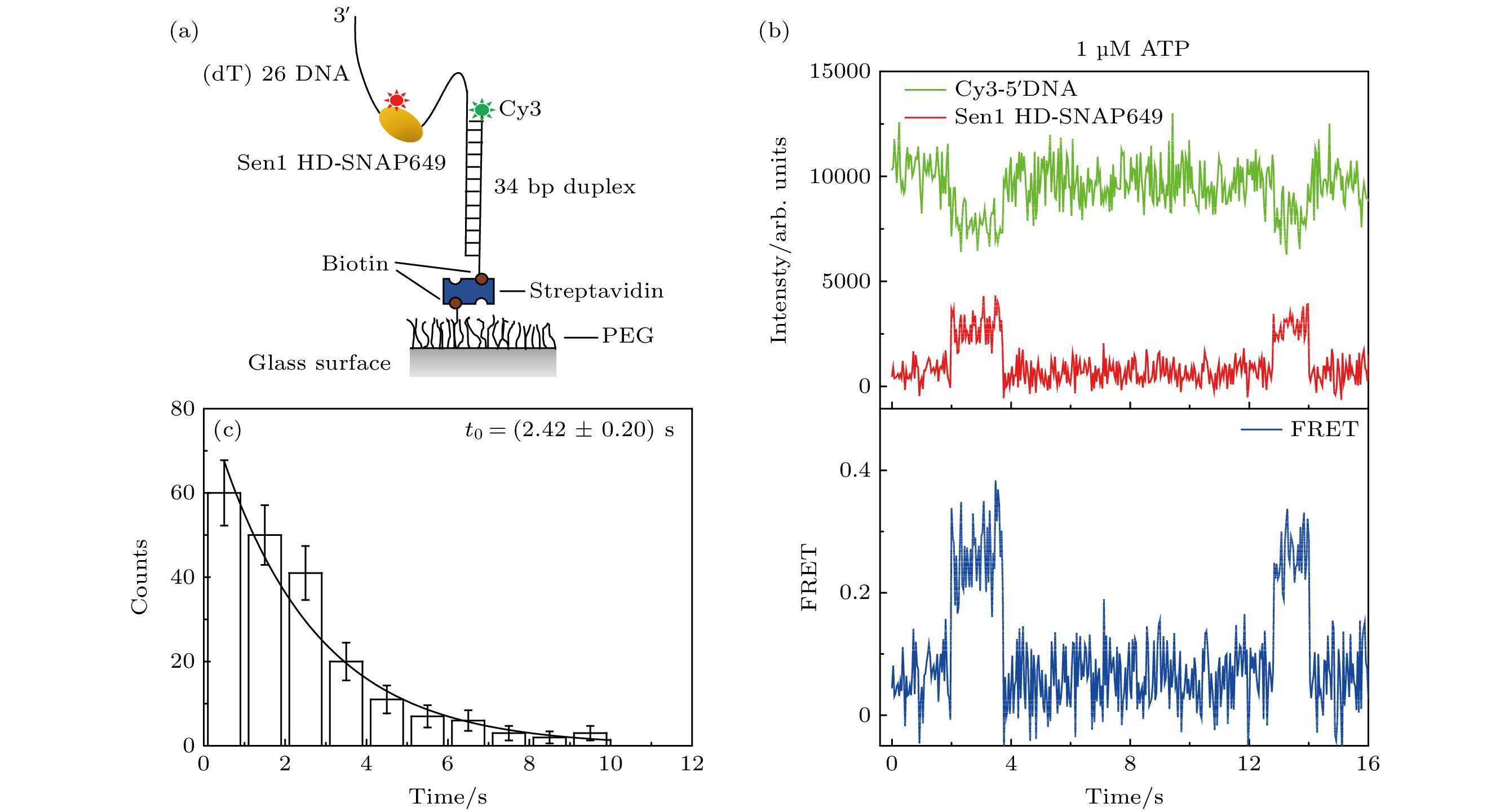
图 3 单分子FRET方法表征Sen1 HD-SNAP649在5'-26ss-34duplex上的行走功能(1 µM ATP条件下) (a) 单分子FRET方法示意图, 在5'-26ss-34duplex结构中, 岔口5'端标有Cy3荧光, Sen1 HD上标有SNAP649荧光; (b) Sen1 HD-SNAP649行走的典型曲线; (c) FRET持续时间的分布图
Figure 3. Sen1 HD-SNAP649 translocation activity on 5'-26ss-34duplex characterized via single-molecule FRET assay under 1 μM ATP condition: (a) Schematic for the assay, in the 5'-26ss-34duplex construct, the 5' end of the fork is labeled with Cy3, and Sen1 HD is labeled with SNAP649; (b) a typical trajectory representing Sen1 HD-SNAP649 translocation on DNA substrate; (c) distribution of the FRET dwell times corresponding to Sen1 HD-SNAP649 translocation.
图 5 单分子FRET方法表征Sen1 HD-SNAP649在3'-26ss-34duplex上的行走(1 µM ATP条件) (a) 实验示意图, 在3'-26ss-34duplex结构中, 岔口3' 端标有Cy3荧光, Sen1 HD上标有SNAP649荧光; (b) Sen1 HD-SNAP649行走的典型曲线; (c) FRET持续时间的分布图
Figure 5. Translocation activity of Sen1 HD-SNAP649 on 3'-26ss-34duplex characterized via single-molecule FRET assay under 1 μM ATP condition: (a) Schematic of the assay, and in the construct of 3'-26ss-34duplex, the 3' end of the fork is labeled with Cy3, and Sen1 HD is labeled with SNAP649; (b) a typical trajectory representing Sen1 HD-SNAP649 translocation; (c) histogram of the FRET dwell times for Sen1 HD-SNAP649 translocation.
表 1 DNA底物序列信息及其应用
Table 1. DNA sequences and their applications.
DNA底物名称 DNA序列(5'-3') 应用 5'-50ss-20duplex 20-nt FAM ssDNA: GTT GGG TAA CGC CAG GGA CG-3'FAM70-nt ssDNA: ATT ACG GAT TCA CTG GCC GTC GTT TTA CAA CGT CGT GAC TGG GAA AAA CGC GTC CCT GGC GTT ACC CAA C DNA退火后形成具有50 nt单链结合位点的20 bp 双链DNA底物, 用于表征Sen1 HD解旋活性 Cy3-48ssDNA AGC TGG ATA CTT ACA GCC ATG GCT GCT GCG AAT ACT CCA TTC CAT CCC 用于验证Sen1 HD与单链DNA的结合能力 5'-26ss-34duplex 5'bio-60ssDNA: 5'Biotin-GCC AGG AGG CTA GCA ACA GTC TTC ATT CAA CCG ACG TCA CAA TAG TGA GTA CCA ATA CCT5'Cy3-34ssDNA: 5'Cy3-TCG GTT GAA TGA AGA CTG TTG CTA GCC TCC TGG C DNA退火后形成具有26 nt 单链结合位点沿5'-3'方向远离DNA岔口的34 bp双链DNA底物, 用于表征Sen1 HD-SNAP649在单链DNA上的行走功能 3'-26ss-34duplex 3'bio-60ssDNA: TCC ATA ACC ATG AGT GAT AAC ACT GCA GCC AAC TTA CTT CTG ACA ACG ATC GGA GGA CCG-3'Biotin3'Cy3-34ssDNA: CGG TCC TCC GAT CGT TGT CAG AAG TAA GTT GGC T-3'Cy3 DNA退火后形成26 nt 单链结合位点沿5'-3'方向靠近DNA岔口的34 bp双链DNA底物, 用于表征Sen1 HD-SNAP649在单链DNA上的行走方向 -
[1] Roeder R G 2019 Nat. Struct. Mol. Biol. 26 783
 Google Scholar
Google Scholar
[2] Jonkers I, Lis J T 2015 Nat. Rev. Mol. Cell. Biol. 16 167
 Google Scholar
Google Scholar
[3] Abbondanzieri E A, Greenleaf W J, Shaevitz J W, Landick R, Block S M 2005 Nature 438 460
 Google Scholar
Google Scholar
[4] You L, Omollo E O, Yu C, Mooney R A, Shi J, Shen L, Wu X, Wen A, He D, Zeng Y, Feng Y, Landick R, Zhang Y 2023 Nature 613 783
 Google Scholar
Google Scholar
[5] Molodtsov V, Wang C, Firlar E, Kaelber J T, Ebright R H 2023 Nature 614 367
 Google Scholar
Google Scholar
[6] West S, Gromak N, Proudfoot N J 2004 Nature 432 522
 Google Scholar
Google Scholar
[7] Porrua O, Libri D 2013 Nat. Struct. Mol. Biol. 20 884
 Google Scholar
Google Scholar
[8] Wang S, Han Z, Libri D, Porrua O, Strick T R 2019 Nat. Commun. 10 1545
 Google Scholar
Google Scholar
[9] Vasiljeva L, Kim M, Mutschler H, Buratowski S, Meinhart A 2008 Nat. Struct. Mol. Biol. 15 795
 Google Scholar
Google Scholar
[10] Arndt K M, Reines D 2015 Annu. Rev. Biochem. 84 381
 Google Scholar
Google Scholar
[11] Rondón A G, Mischo H E, Kawauchi J, Proudfoot N J 2009 Mol. Cell. 36 88
 Google Scholar
Google Scholar
[12] Jia X, Li Y, Wang T, Bi L, Guo L, Chen Z, Zhang X, Ye S, Chen J, Yang B, Sun B 2023 Embo. J. 42 e111703
 Google Scholar
Google Scholar
[13] Saper G, Hess H 2020 Chem. Rev. 120 288
 Google Scholar
Google Scholar
[14] Nishizaka T 2010 Adv. Biochem. Eng. Biotechnol. 119 3
 Google Scholar
Google Scholar
[15] Martin-Tumasz S, Brow D A 2015 J. Biol. Chem. 290 22880
 Google Scholar
Google Scholar
[16] Han Z, Libri D, Porrua O 2017 Nucleic. Acids. Res. 45 1355
 Google Scholar
Google Scholar
[17] Skourti-Stathaki K, Proudfoot N J, Gromak N 2011 Mol. Cell. 42 794
 Google Scholar
Google Scholar
[18] Ha T, Enderle T, Ogletree D F, Chemla D S, Selvin P R, Weiss S 1996 Proc. Natl. Acad. Sci. USA 93 6264
 Google Scholar
Google Scholar
[19] Maki A H, Co T 1976 Biochemistry 15 1229
 Google Scholar
Google Scholar
[20] Leonaitė B, Han Z, Basquin J, Bonneau F, Libri D, Porrua O, Conti E 2017 Embo. J. 36 1590
 Google Scholar
Google Scholar
[21] Shi J, Wang F, Li F, Wang L, Xiong Y, Wen A, Jin Y, Jin S, Gao F, Feng Z, Li J, Zhang Y, Shang Z, Wang S, Feng Y, Lin W 2022 Nucleic. Acids. Res. 50 5974
 Google Scholar
Google Scholar
[22] Fiorini F, Bagchi D, Le Hir H, Croquette V 2015 Nat. Commun. 6 7581
 Google Scholar
Google Scholar
[23] Chakrabarti S, Jayachandran U, Bonneau F, Fiorini F, Basquin C, Domcke S, Le Hir H, Conti E 2011 Mol. Cell. 41 693
 Google Scholar
Google Scholar
[24] Hazelbaker D Z, Marquardt S, Wlotzka W, Buratowski S 2013 Mol. Cell. 49 55
 Google Scholar
Google Scholar
Catalog
Metrics
- Abstract views: 6708
- PDF Downloads: 122
- Cited By: 0



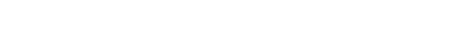









 DownLoad:
DownLoad: